This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison, Genetics 564
What is protein homology?
Homologous proteins are proteins form two different species that derived form a common ancestor. As the two species evolved, their proteins sequences will deviate to different degrees depending on a variety of facors. Determining that two proteins are homologous may help researchers learn more about their protein of interest or perform test in model organisms.
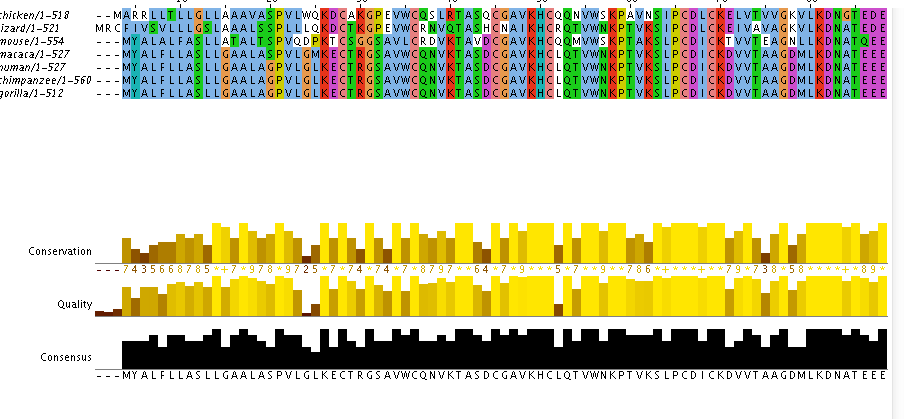
How well conserved in the PSAP protein?
What is protein phylogeny?
Protein phylogeny is a way of organizing data on the protein sequence for particular gene from a variety of organisms. After aligning protein sequences, phylogenic trees are created which give relationships on the similarity between a variety of sequences. There are several different methods of generating these trees. A few of the methods are listed bellow.
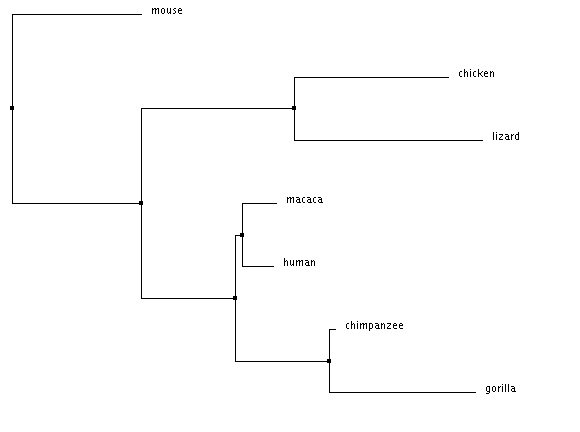
The protein sequences were aligned using Clustal Omega. The trees bellow were calculated using the alignment seen above.
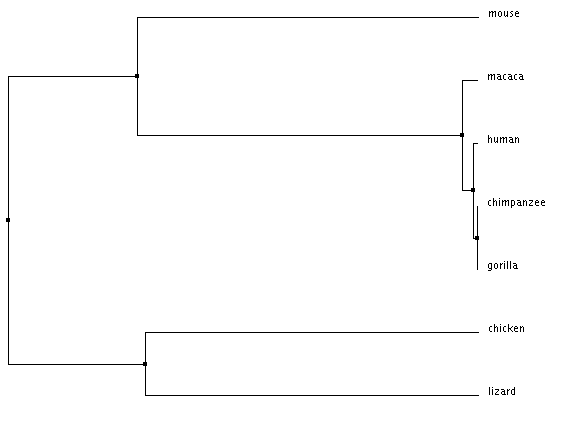
Average distance tree using % identity
What is a neighbor joining tree?
Neighbor joining trees account for difference between species relatedness using the branch length. The longer a the branch length, the further apart to the two protein sequences are from each other.
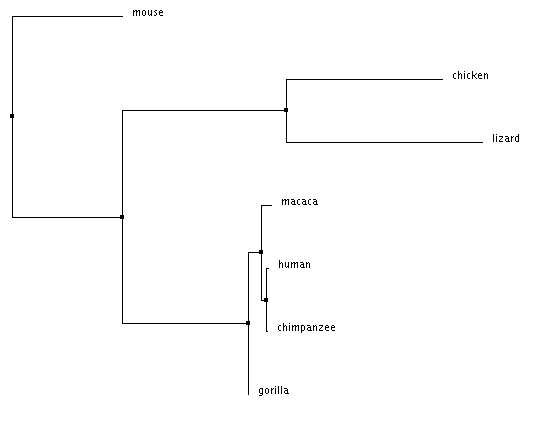
Neighbor joining tree using % identity
What is the BLOSUM scoring matrix?
The BLOSUM matrix is a standardized way of assigning values on the differences between amino acids. Because the chemical composition of each amino acid is unique, they all function in different ways too. However, amino acids with similar chemical structure often function in similar ways. Therefore, not all substitutions in the amino acid sequence are equal. The numerical values assigned to amino acid changes can be found here.
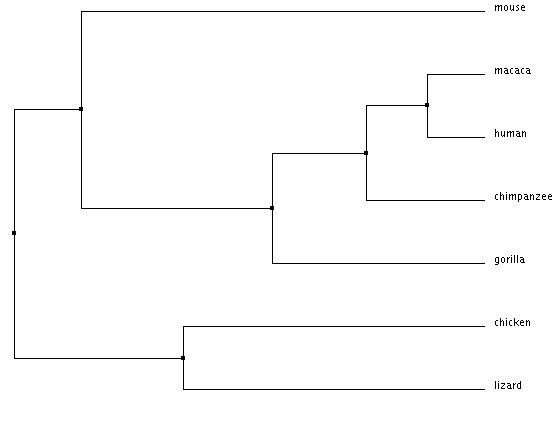
Average distance tree using BLOSUM62
Neighbor joining tree using BLOSUM62
What are protein domains?
Protein domains are conserved sequences of amino acids that have specific tertiary structures. They often give a protein its specific function or property based on the structure of the domain. The role of domains can vary widely from binding to specific molecules or cleaving other molecules.
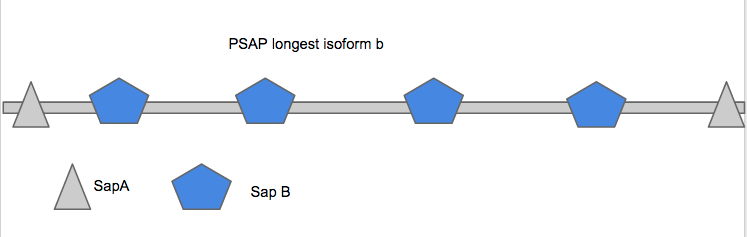
Protein Domains of PSAP
PSAP encodes for a precursor protein that is cleaved into four products: saposins A, B, C, and D. The cleavage occurs inside the lysosome of the cell. The longest isoform of PSAP is isoform b and it is 537 amino acids long.
In PSAP, there are only two protein domains recognized by the websites Pfam and SMART. The domains recognized are SapA and SapB. The first SapA is located between the 21st and 54th amino acid and the second is located from 494th to the 527th amino acid. The two SapA domains will be removed during the cleavage of PSAP and the four SapB domains will become the active saposin products: A, B, C, and D [1].
Interestingly, the Sap B domains contain six cystine residues that are highly conserved and form three disulfide bridges [2].
In PSAP, there are only two protein domains recognized by the websites Pfam and SMART. The domains recognized are SapA and SapB. The first SapA is located between the 21st and 54th amino acid and the second is located from 494th to the 527th amino acid. The two SapA domains will be removed during the cleavage of PSAP and the four SapB domains will become the active saposin products: A, B, C, and D [1].
Interestingly, the Sap B domains contain six cystine residues that are highly conserved and form three disulfide bridges [2].
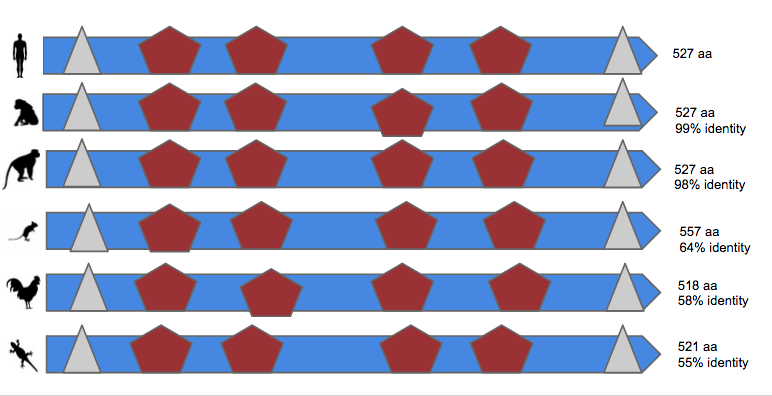
How well conserved are the protein domains?
What are protein interaction networks?
Protein interaction networks show all of the proteins that a protein of interest interacts with and how those proteins interact with each other. The networks are generated from databases were researchers will annotate how proteins interact using model organisms. These networks can be used to anylize webs of interest and uncvover unecpected interactions or patterns of interaction. This information can be useful when attempting to better understand a cellular process or protein.
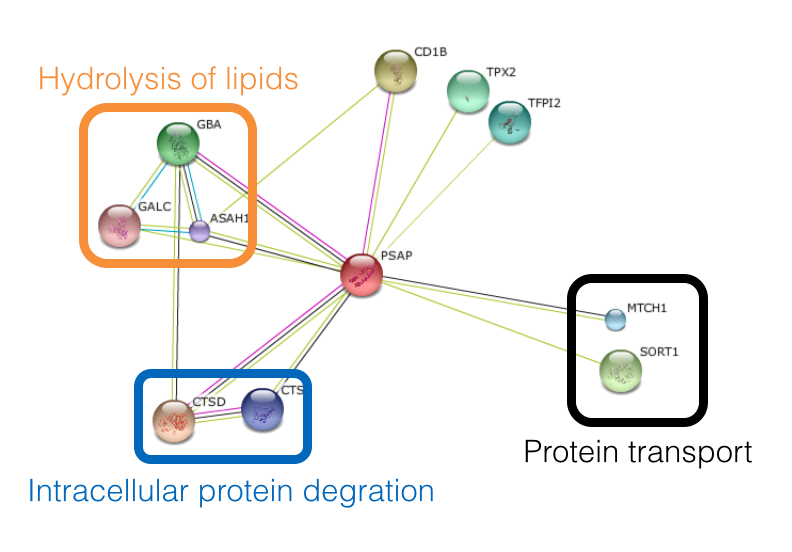
Using the String Database, this protein interaction network was formed. The majority of the proteins that PSAP interacts with can be divided into three different groups: protein transport, intracellular protein degradation, and lipid hydrolysis.
Post transnational modifications
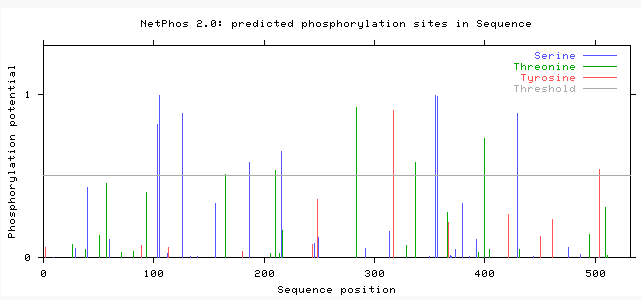
After a protein is translated, modifications can still be made to the amino acids. These modification can be vary from how a protein folds to phosphorylation of an amino acids. Using the NetPhos 2.0 server, phosphorylation sites on the PSAP protein were predicted on serine, threonine, and tyrosine. The database predicted the phosphorylation of 8 serines, 5 threonines. and 2 tryosines. The graph bellow shows the results.
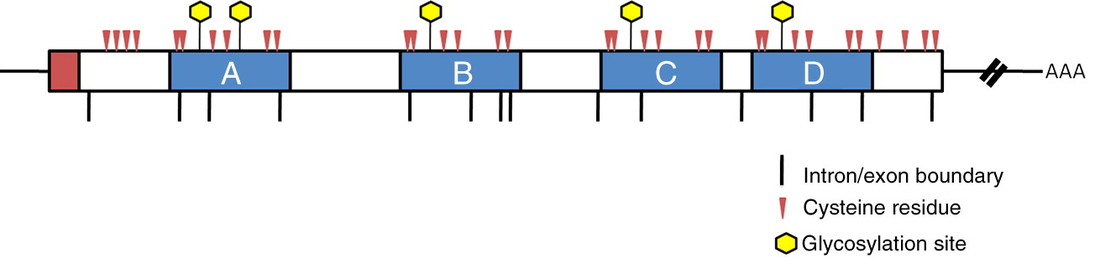
Previous studies have also identified that the PSAP protein get glycosylated in the Golgi and are essential in transportation of the protein to the lysosome [3]. The glycosylation sites can be seen in the image bellow (image taken from Tamargo et al., The role of saposin C in Gaucher disease).
Citations
1. Ponting CP (1994). Acid sphingomyelinase possesses a domain homologous to its activator proteins: saposins B and D, Protein Sci., 3 (2), 359–361.
2. Vaccaro R, Salvioli M, Tatti, Ciaffoni, F. 1999. Saposins and their interaction with lipids, Neurochem. Res., 24, 307–314.
3. Lefrancois, S, et al. 2003. The lysosomal trafficking of sphingolipid activator proteins (SAPs) is mediated by sortilin, The EMBO Journal 22, 6430-6437.
Media
Chimpanzee: http://www.clipartbest.com/chimpanzee-clip-art
Chicken: http://www.clipartpanda.com/categories/cooked-chicken-clipart
Mouse: http://www.clker.com/clipart-simple-cartoon-mouse-2.html
Gorilla: http://photolabels.co/animals/628966/gorilla-cartoon-character-mascot-design/
Cat: http://www.clipartpanda.com/categories/cat-20clip-20art
Macaque: http://www.clipartpanda.com/categories/cat-20clip-20art
Lizard: http://www.clipartpanda.com/categories/cute-lizard-clipart
Cod: http://www.clipartsfree.net/medium/23277-cod-fish-clipart.html
1. Ponting CP (1994). Acid sphingomyelinase possesses a domain homologous to its activator proteins: saposins B and D, Protein Sci., 3 (2), 359–361.
2. Vaccaro R, Salvioli M, Tatti, Ciaffoni, F. 1999. Saposins and their interaction with lipids, Neurochem. Res., 24, 307–314.
3. Lefrancois, S, et al. 2003. The lysosomal trafficking of sphingolipid activator proteins (SAPs) is mediated by sortilin, The EMBO Journal 22, 6430-6437.
Media
Chimpanzee: http://www.clipartbest.com/chimpanzee-clip-art
Chicken: http://www.clipartpanda.com/categories/cooked-chicken-clipart
Mouse: http://www.clker.com/clipart-simple-cartoon-mouse-2.html
Gorilla: http://photolabels.co/animals/628966/gorilla-cartoon-character-mascot-design/
Cat: http://www.clipartpanda.com/categories/cat-20clip-20art
Macaque: http://www.clipartpanda.com/categories/cat-20clip-20art
Lizard: http://www.clipartpanda.com/categories/cute-lizard-clipart
Cod: http://www.clipartsfree.net/medium/23277-cod-fish-clipart.html
Mitchell Coplan, [email protected], 5/10/15, gen564